# Import the following libraries
# For fetching from the Raster API
import requests
# For making maps
import folium
import folium.plugins
from folium import Map, TileLayer
# For talking to the STAC API
from pystac_client import Client
# For working with data
import pandas as pd
# For making time series
import matplotlib.pyplot as plt
# For formatting date/time data
import datetime
# Custom functions for working with GHGC data via the API
import ghgc_utilsMiCASA Land Carbon Flux
Access this Notebook
You can launch this notebook in the US GHG Center JupyterHub (requires access) by clicking the following link: MiCASA Land Carbon Flux. If you are a new user, you should first sign up for the hub by filling out this request form and providing the required information.
If you do not have a US GHG Center Jupyterhub account, you can access this notebook through MyBinder by clicking the button below.
Table of Contents
Data Summary and Application
- Spatial coverage: Global
- Spatial resolution: 0.1° x 0.1°
- Temporal extent: January 1, 2001 - December 31, 2023
- Temporal resolution: Daily and Monthly Averages
- Unit: Grams of Carbon per square meter per day (g Carbon/m²/day)
- Utility: Climate Research
For more information, visit the MiCASA Land Carbon Flux data overview page.
Approach
- Identify available dates and temporal frequency of observations for a given collection using the US Greenhouse Gas Center (GHGC) Application Programming Interface (API)
/stacendpoint. The collection processed in this notebook is the MiCASA Land Carbon Flux data product - Pass the STAC item into the raster API
/collections/{collection_id}/items/{item_id}/{tile_matrix_set_id}/tilejson.jsonendpoint - Using
folium.plugins.DualMap, visualize two tiles (side-by-side), allowing time point comparison - After the visualization, perform zonal statistics for a given polygon
About the Data
MiCASA Land Carbon Flux
This dataset presents a variety of carbon flux parameters derived from the Más Informada Carnegie-Ames-Stanford-Approach (MiCASA) model. The model’s input data includes air temperature, precipitation, incident solar radiation, a soil classification map, and several satellite-derived products. All model calculations are driven by analyzed meteorological data from NASA’s Modern-Era Retrospective analysis for Research and Application, Version 2 (MERRA-2). The resulting product provides global, daily data at 0.1 degree resolution from January 2001 through December 2023. It includes carbon flux variables expressed in units of kilograms of carbon per square meter per day (kg Carbon/m²/day) from net primary production (NPP), heterotrophic respiration (Rh), wildfire emissions (FIRE), fuel wood burning emissions (FUEL), net ecosystem exchange (NEE), and net biosphere exchange (NBE). The latter two are derived from the first four (see Scientific Details below). MiCASA is an extensive revision of the CASA – Global Fire Emissions Database, version 3 (CASA-GFED3) product. CASA-GFED3 and earlier versions of MERRA-driven CASA-GFED carbon fluxes have been used in several atmospheric carbon dioxide (CO₂) transport studies, serve as a community standard for priors of flux inversion systems, and through the support of NASA’s Carbon Monitoring System (CMS), help characterize, quantify, understand and predict the evolution of global carbon sources and sinks.
For more information regarding this dataset, please visit the MiCASA Land Carbon Flux data overview page.
Terminology
Navigating data via the GHGC API, you will encounter terminology that is different from browsing in a typical filesystem. We’ll define some terms here which are used throughout this notebook.
catalog: All datasets available at the/stacendpointcollection: A specific dataset, e.g. MiCASA Land Carbon Fluxitem: One data file (i.e. granule) in the dataset, e.g. one daily file of carbon fluxesasset: A variable available within the granule, e.g. net primary productivity or net ecosystem exchangeSTAC API: SpatioTemporal Asset Catalogs - Endpoint for fetching metadata about available datasetsRaster API: Endpoint for fetching data itself, for imagery and statistics
Install the Required Libraries
Required libraries are pre-installed on the US GHG Center Hub. If you need to run this notebook elsewhere, please install them with this line in a code cell:
%pip install requests folium rasterstats pystac_client pandas matplotlib –quiet
Query the STAC API
STAC API Collection Names
Now, you must fetch the dataset from the STAC API by defining its associated STAC API collection ID as a variable. The collection ID, also known as the collection name, for the MiCASA Land Carbon Flux dataset is micasa-carbonflux-monthgrid-v1.
# Provide the STAC and RASTER API endpoints
# The endpoint is referring to a location within the API that executes a request on a data collection nesting on the server.
# The STAC API is a catalog of all the existing data collections that are stored in the GHG Center.
STAC_API_URL = "https://earth.gov/ghgcenter/api/stac"
# The RASTER API is used to fetch collections for visualization
RASTER_API_URL = "https://earth.gov/ghgcenter/api/raster"
# The collection name is used to fetch metadata from the STAC API.
collection_name = "micasa-carbonflux-monthgrid-v1"# Fetch the collection from the STAC API using the appropriate endpoint
# The 'pystac_client' library makes an HTTP request
catalog = Client.open(STAC_API_URL)
collection = catalog.get_collection(collection_name)
# Print the properties of the collection to the console
collectionExamining the contents of our collection under the temporal variable, we see that the data is available from January 2001 to December 2023. By looking at the dashboard:time density, we observe that the periodic frequency of these observations is monthly.
#items = list(collection.get_items()) # Convert the iterator to a list
#print(f"Found {len(items)} items")# The search function can be used to find data within a specific time frame
search = catalog.search(
collections=collection_name,
datetime=['2010-01-01T00:00:00Z','2010-12-31T00:00:00Z']
)
# Take a look at the items we found
for item in search.item_collection():
print(item)<Item id=micasa-carbonflux-monthgrid-v1-201012>
<Item id=micasa-carbonflux-monthgrid-v1-201011>
<Item id=micasa-carbonflux-monthgrid-v1-201010>
<Item id=micasa-carbonflux-monthgrid-v1-201009>
<Item id=micasa-carbonflux-monthgrid-v1-201008>
<Item id=micasa-carbonflux-monthgrid-v1-201007>
<Item id=micasa-carbonflux-monthgrid-v1-201006>
<Item id=micasa-carbonflux-monthgrid-v1-201005>
<Item id=micasa-carbonflux-monthgrid-v1-201004>
<Item id=micasa-carbonflux-monthgrid-v1-201003>
<Item id=micasa-carbonflux-monthgrid-v1-201002>
<Item id=micasa-carbonflux-monthgrid-v1-201001># Examine the first item in the search results
# Keep in mind that a list starts from 0, 1, 2... therefore items[0] is referring to the first item in the list/collection
items = search.item_collection()
items[0]# Restructure our items into a dictionary where keys are the datetime items
# Then we can query more easily by date/time, e.g. "2020-05"
items_dict = {item.properties["datetime"][:7]: item for item in items}# Before we go further, let's pick which asset to focus on for the remainder of the notebook.
# We'll focus on net primary productivity, so our asset of interest is:
asset_name = "npp"Creating Maps Using Folium
You will now explore changes in the land atmosphere Carbon flux Net Primary Productivity at a given location. You will visualize the outputs on a map using folium.
Fetch Imagery Using Raster API
Here we get information from the Raster API which we will add to our map in the next section.
# Specify two date/times that you would like to visualize, using the format of items_dict.keys()
dates = ["2010-01","2010-07"]Below, we use some statistics of the raster data to set upper and lower limits for our color bar. These are saved as rescale_values, and will be passed to the Raster API in the following step(s).
# Extract collection name and item ID for the first date
observation_date_1 = items_dict[dates[0]]
collection_id = observation_date_1.collection_id
item_id = observation_date_1.id
tile_matrix_set_id = "WebMercatorQuad"
# Select relevant asset (NPP)
object = observation_date_1.assets[asset_name]
raster_bands = object.extra_fields.get("raster:bands", [{}])
# Print raster bands' information
raster_bands[{'unit': 'g C m-2 day-1',
'scale': 1.0,
'nodata': 'nan',
'offset': 0.0,
'sampling': 'area',
'data_type': 'float32',
'histogram': {'max': 5.948176860809326,
'min': -0.3724845349788666,
'count': 11,
'buckets': [488513, 9595, 5022, 3321, 2776, 3213, 4254, 4777, 2672, 145]},
'statistics': {'mean': 0.1637769341468811,
'stddev': 0.7254217411478966,
'maximum': 5.948176860809326,
'minimum': -0.3724845349788666,
'valid_percent': 100.0}}]# Use raster band statistics to generate an appropriate color bar range.
rescale_values = {
"max": raster_bands[0].get("histogram", {}).get("max"),
"min": raster_bands[0].get("histogram", {}).get("min"),
}
print(rescale_values){'max': 5.948176860809326, 'min': -0.3724845349788666}Now, you will pass the item id, collection name, asset name, and the rescale values to the Raster API endpoint, along with a colormap. This step is done twice, one for each date/time you will visualize, and tells the Raster API which collection, item, and asset you want to view, specifying the colormap and colorbar ranges to use for visualization. The API returns a JSON with information about the requested image. Each image will be referred to as a tile.
# Choose a colormap for displaying the data
# Make sure to capitalize per Matplotlib standard colormap names
# For more information on Colormaps in Matplotlib, please visit https://matplotlib.org/stable/users/explain/colors/colormaps.html
color_map = "PuRd"# Make a GET request to retrieve information for your first date/time
observation_date_1_tile = requests.get(
f"{RASTER_API_URL}/collections/{collection_id}/items/{item_id}/{tile_matrix_set_id}/tilejson.json?"
f"&assets={asset_name}"
f"&color_formula=gamma+r+1.05&colormap_name={color_map.lower()}"
f"&rescale={rescale_values['min']},{rescale_values['max']}"
).json()
# Print the properties of the retrieved granule to the console
observation_date_1_tile{'tilejson': '2.2.0',
'version': '1.0.0',
'scheme': 'xyz',
'tiles': ['https://earth.gov/ghgcenter/api/raster/collections/micasa-carbonflux-monthgrid-v1/items/micasa-carbonflux-monthgrid-v1-201001/tiles/WebMercatorQuad/{z}/{x}/{y}@1x?assets=npp&color_formula=gamma+r+1.05&colormap_name=purd&rescale=-0.3724845349788666%2C5.948176860809326'],
'minzoom': 0,
'maxzoom': 24,
'bounds': [-180.0, -90.0, 179.99999999999994, 90.0],
'center': [-2.842170943040401e-14, 0.0, 0]}# Repeat the above for your second date/time
# Note that we do not calculate new rescale_values for this tile -
# We want date tiles 1 and 2 to have the same colorbar range for visual comparison.
observation_date_2 = items_dict[dates[1]]
# Extract collection name and item ID
collection_id = observation_date_2.collection_id
item_id = observation_date_2.id
# Make a GET request to retrieve tile information
observation_date_2_tile = requests.get(
f"{RASTER_API_URL}/collections/{collection_id}/items/{item_id}/{tile_matrix_set_id}/tilejson.json?"
f"&assets={asset_name}"
f"&color_formula=gamma+r+1.05&colormap_name={color_map.lower()}"
f"&rescale={rescale_values['min']},{rescale_values['max']}"
).json()
# Print the properties of the retrieved granule to the console
observation_date_2_tile{'tilejson': '2.2.0',
'version': '1.0.0',
'scheme': 'xyz',
'tiles': ['https://earth.gov/ghgcenter/api/raster/collections/micasa-carbonflux-monthgrid-v1/items/micasa-carbonflux-monthgrid-v1-201007/tiles/WebMercatorQuad/{z}/{x}/{y}@1x?assets=npp&color_formula=gamma+r+1.05&colormap_name=purd&rescale=-0.3724845349788666%2C5.948176860809326'],
'minzoom': 0,
'maxzoom': 24,
'bounds': [-180.0, -90.0, 179.99999999999994, 90.0],
'center': [-2.842170943040401e-14, 0.0, 0]}Generate Map
For this example, we’ll look at NPP over the Amazon Rainforest, South America.
First, let’s determine an area of interest (AOI) to visualize via GeoJSON.
# Set a name for the AOI to use in plots later
aoi_name = "Amazon Rainforest"
# The Area of Interest (AOI) is set to the Amazon Rainforest, South America
aoi = {
"type": "Feature",
"properties": {},
"geometry": {
"coordinates": [
[
# [longitude, latitude]
[-74.0, -3.0], # Southwest Bounding Coordinate
[-74.0, 5.0], # Northwest Bounding Coordinate
[-60.0, 5.0], # Northeast Bounding Coordinate
[-60.0, -3.0], # Southeast Bounding Coordinate
[-74.0, -3.0] # Closing the polygon at the Southwest Bounding Coordinate
]
],
"type": "Polygon",
},
}# Initialize the map, specifying the center of the map and the starting zoom level.
# 'folium.plugins' allows mapping side-by-side via 'DualMap'
# Map is centered on the position specified by "location=(lat,lon)"
map_ = folium.plugins.DualMap(location=(0, -66), zoom_start=5)
# Define the first map layer using the tile fetched for the first date
# The TileLayer library helps in manipulating and displaying raster layers on a map
map_layer_observation_date_1 = TileLayer(
tiles=observation_date_1_tile["tiles"][0], # Path to retrieve the tile
attr="US GHG Center", # Set the attribution
opacity=0.8, # Adjust the transparency of the layer
name=f"{dates[0]} NPP", # Title for the layer
overlay= True, # The layer can be overlaid on the map
legendEnabled = True # Enable displaying the legend on the map
)
# Add the first layer to the Dual Map
# This will appear on the left side, specified by 'm1'
map_layer_observation_date_1.add_to(map_.m1)
# Define the second map layer using the tile fetched for the second date
map_layer_observation_date_2 = TileLayer(
tiles=observation_date_2_tile["tiles"][0], # Path to retrieve the tile
attr="US GHG Center", # Set the attribution
opacity=0.8, # Adjust the transparency of the layer
name=f"{dates[1]} NPP", # Title for the layer
overlay= True, # The layer can be overlaid on the map
legendEnabled = True # Enable displaying the legend on the map
)
# Add the second layer to the Dual Map
# This will appear on the left side, specified by 'm2'
map_layer_observation_date_2.add_to(map_.m2)
# Display data marker on each map
folium.Marker((0, -66), tooltip=aoi_name).add_to(map_)
# Display AOI on each map
folium.GeoJson(aoi, name=f"{aoi_name} AOI",style_function=lambda feature: {"fillColor": "none"}).add_to(map_)
# Add a layer control to switch between map layers
folium.LayerControl(collapsed=False).add_to(map_)
# Add colorbar
# We can use 'generate_html_colorbar' from the 'ghgc_utils' module
# to create an HTML colorbar representation.
legend_html = ghgc_utils.generate_html_colorbar(color_map,rescale_values,label='NPP (g Carbon/m2/daily)',dark=True)
# Add colorbar to the map
map_.get_root().html.add_child(folium.Element(legend_html))
# Visualize the Dual Map
map_Calculate Zonal Statistics
We will perform zonal statistics and create a dataframe using our previously defined AOI for the Amazon Rainforest, the items from our initial catalog search spanning from January 1, 2010 through December 31, 2010, and the “npp” asset.
We’ll generate statistics for the data in our AOI using a function from the ghgc_utils module, which fetches the data and its statistics from the Raster API.
%%time
# %%time = Wall time (execution time) for running the code below
# Statistics will be returned as a Pandas DataFrame
df = ghgc_utils.generate_stats(items,aoi,url=RASTER_API_URL,asset=asset_name)
# Print the first five rows of statistics from our DataFrame
df.head(5)Generating stats...
Done!
CPU times: total: 2.05 s
Wall time: 9.25 s| datetime | min | max | mean | count | sum | std | median | majority | minority | unique | histogram | valid_percent | masked_pixels | valid_pixels | percentile_2 | percentile_98 | date | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 2010-12-01T00:00:00Z | 0.00000000000000000000 | 4.89262914657592773438 | 4.21476507186889648438 | 11200.00000000000000000000 | 47205.36718750000000000000 | 0.67433501265570172656 | 4.46684360504150390625 | 0.00000000000000000000 | 0.09475493431091308594 | 11151.00000000000000000000 | [[17, 11, 43, 129, 196, 517, 399, 617, 2585, 6... | 100.00000000000000000000 | 0.00000000000000000000 | 11200.00000000000000000000 | 2.06423521041870117188 | 4.74646377563476562500 | 2010-12-01 00:00:00+00:00 |
| 1 | 2010-11-01T00:00:00Z | 0.00000000000000000000 | 4.99066686630249023438 | 4.32144927978515625000 | 11200.00000000000000000000 | 48400.23437500000000000000 | 0.67623456182856067631 | 4.56732463836669921875 | 0.00000000000000000000 | 0.11350738257169723511 | 11152.00000000000000000000 | [[21, 13, 38, 124, 154, 462, 442, 634, 2499, 6... | 100.00000000000000000000 | 0.00000000000000000000 | 11200.00000000000000000000 | 2.11893200874328613281 | 4.85591602325439453125 | 2010-11-01 00:00:00+00:00 |
| 2 | 2010-10-01T00:00:00Z | 0.00000000000000000000 | 5.11889410018920898438 | 4.37894201278686523438 | 11200.00000000000000000000 | 49044.14843750000000000000 | 0.70461959934651896553 | 4.64984226226806640625 | 0.00000000000000000000 | 0.00312864035367965698 | 11153.00000000000000000000 | [[21, 14, 48, 137, 155, 492, 540, 620, 2731, 6... | 100.00000000000000000000 | 0.00000000000000000000 | 11200.00000000000000000000 | 2.05418276786804199219 | 4.93095159530639648438 | 2010-10-01 00:00:00+00:00 |
| 3 | 2010-09-01T00:00:00Z | 0.00000000000000000000 | 5.20381402969360351562 | 4.25992488861083984375 | 11200.00000000000000000000 | 47711.16015625000000000000 | 0.64925805993084317880 | 4.48059511184692382812 | 0.00000000000000000000 | 0.07202455401420593262 | 11170.00000000000000000000 | [[18, 13, 42, 135, 184, 544, 540, 827, 7751, 1... | 100.00000000000000000000 | 0.00000000000000000000 | 11200.00000000000000000000 | 2.13949847221374511719 | 4.92846536636352539062 | 2010-09-01 00:00:00+00:00 |
| 4 | 2010-08-01T00:00:00Z | 0.00000000000000000000 | 5.27340316772460937500 | 4.15344047546386718750 | 11200.00000000000000000000 | 46518.53515625000000000000 | 0.59038417891895100809 | 4.29437303543090820312 | 0.00000000000000000000 | 0.00740983802825212479 | 11165.00000000000000000000 | [[15, 13, 39, 106, 164, 576, 668, 2322, 6589, ... | 100.00000000000000000000 | 0.00000000000000000000 | 11200.00000000000000000000 | 2.30402040481567382812 | 4.91697549819946289062 | 2010-08-01 00:00:00+00:00 |
Time-Series Analysis
You can now explore the net primary production values in 2010 for the Amazon Rainforest in South America. You can plot the data set using the code below:
# Determine the width and height of the plot using the 'matplotlib' library
# Figure size: 20 representing the width, 10 representing the height
fig = plt.figure(figsize=(10, 5))
# Change 'which_stat' below if you would rather look at a different statistic, like minimum or maximum.
which_stat = "mean"
# Plot the time series analysis of the daily NPP Values for Amazon Rainforest, South America
plt.plot(
df["date"], # X-axis: date
df[which_stat], # Y-axis: NPP value
color="#BA4066", # Line color in hex format
linestyle="-", # Line style
linewidth=2, # Line width
label=asset_name.upper(), # Legend label
)
# Display legend
plt.legend()
# Insert label for the X-axis
plt.xlabel("Years")
# Insert label for the Y-axis
plt.ylabel("g Carbon/m2/day")
# Insert title for the plot
plt.title(f"{which_stat.capitalize()} {asset_name.upper()} for the {aoi_name}")
# Add data citation
plt.text(
df["date"].iloc[-1], # X-coordinate of the text
df[which_stat].min(), # Y-coordinate of the text
# Text to be displayed
f"Source: {collection.title}",
fontsize=8, # Font size
horizontalalignment="left", # Horizontal alignment
verticalalignment="top", # Vertical alignment
color="blue", # Text color
)
plt.show()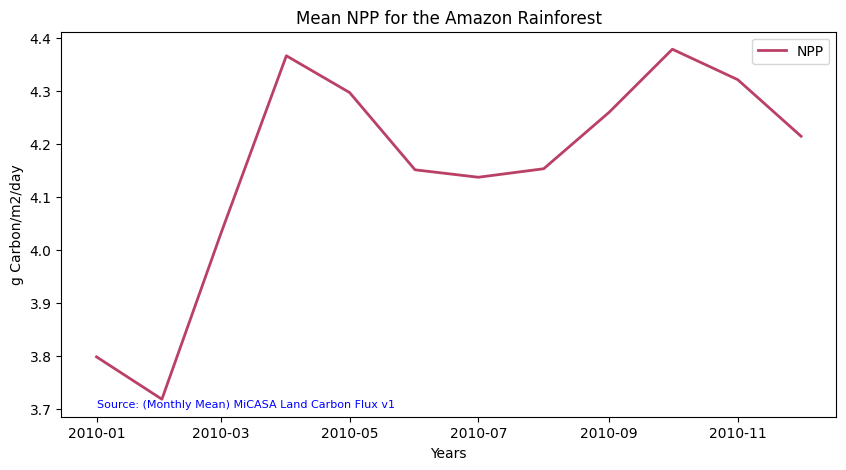
Summary
In this notebook, we have successfully completed the following steps for the MiCASA Land Carbon Flux dataset:
- Install and import the necessary libraries
- Fetch the collection from STAC using the appropriate endpoints
- Count the number of existing granules within the collection
- Map and compare the Net Primary Production (NPP) levels over the Amazon Rainforest, South America area for two distinctive months
- Create a table that displays the minimum, maximum, and sum of the Net Primary Production (NPP) values for a specified region
- Generate a time-series graph of the Net Primary Production (NPP) values for a specified region
If you have any questions regarding this user notebook, please contact us using the feedback form.